FT-IR, NBO, MEP, NLO and Molecular Docking Studies of 8-Epi-9-Deoxy Goniopypyrone
Sreelekshmi PB, Harikumar B, Resmi KS, Sheena Mary Y, Yohannan Panicker C, Van Alsenoy C
Sreelekshmi PB1, Harikumar B1, Resmi KS2, Sheena Mary Y3, Yohannan Panicker C3, Van Alsenoy C4
Department of Chemistry, TKM College of Arts and Science, Kollam, Kerala, India
Department of Physics, TKM College of Arts and Science, Kollam, Kerala, India
Department of Physics, Fatima Mata National College, Kollam, Kerala, India
Chemistry Department, University of Antwerp, Antwerp, Belgium
- *Corresponding Author:
- Yohannan Panicker
C Department of Physics, Fatima Mata National College
Kollam, Kerala
India
E-mail: cyphyp@rediffmail.com
Received date: September 17, 2015; Accepted date: October 01, 2015; Published date: October 05, 2015
Citation: Sreelekshmi PB, Harikumar B, Resmi KS, et al. FT-IR, NBO, MEP, NLO and Molecular Docking Studies of 8-Epi-9-Deoxy Goniopypyrone. Chem Inform. 2015, 1:2.
Abstract
FT-IR spectrum of 8-epi-9-deoxygoniopypyrone (IUPAC name: 8-hydroxy-7-phenyl-2,6-dioxa bicyclo[3.3.1]nonan-3-one) was recorded and analyzed. The vibrational wave numbers were computed using HF and DFT method and are assigned with the help of potential energy distribution method. The stability of the molecule arising from hyper conjugative interaction and charge delocalization has been analyzed using NBO analysis. The HOMO and LUMO analysis are used to determine the charge transfer within the molecule. From the MEP analysis, it is evident that the negative charge covers the C=O group and positive region is over the hydroxyl group. The molecular docking study suggests that the compound might exhibit inhibitory activity against human mutant p53.
Keywords
DFT; Pyrone; FT-IR; Molecular docking; MEP; NBO; NLO
Introduction
Goniothalamus is an important genus of annonaceae family which is rich in highly bioactive lactones and acetogenins [1-6]. Goniothalamus wightii Hook.f and Thoms is a small sized aromatic tree belonging to annonaceae family, growing in the Peppara Wild Life Sanctury of the Nedumangadu forest region of Bonacadu Hills, Trivandrum, Kerala, India [7]. As part of our ongoing phytochemical investigation, we have isolated 8-epi- 9-deoxygoniopypyrone. The structure and three dimensional packing of 8-epi-9-deoxygoniopypyrone was reported [8,9]. In the present work, the IR spectrum of the title compound is reported with the hope that the results of the present work would be helpful in future synthesis of more potent derivatives. The energies, degrees of hybridization, population of lone pairs of oxygen, energies of their interaction with the anti-bonding orbital of the rings and the electron density distributions and stabilization energies have been calculated by natural bond orbital analysis to predict clear evidence of stabilization originating from the hyper conjugation of various intra-molecular interactions. The molecular docking studies are also reported to the potent biological properties of the title compound.
Plant material and experimental details
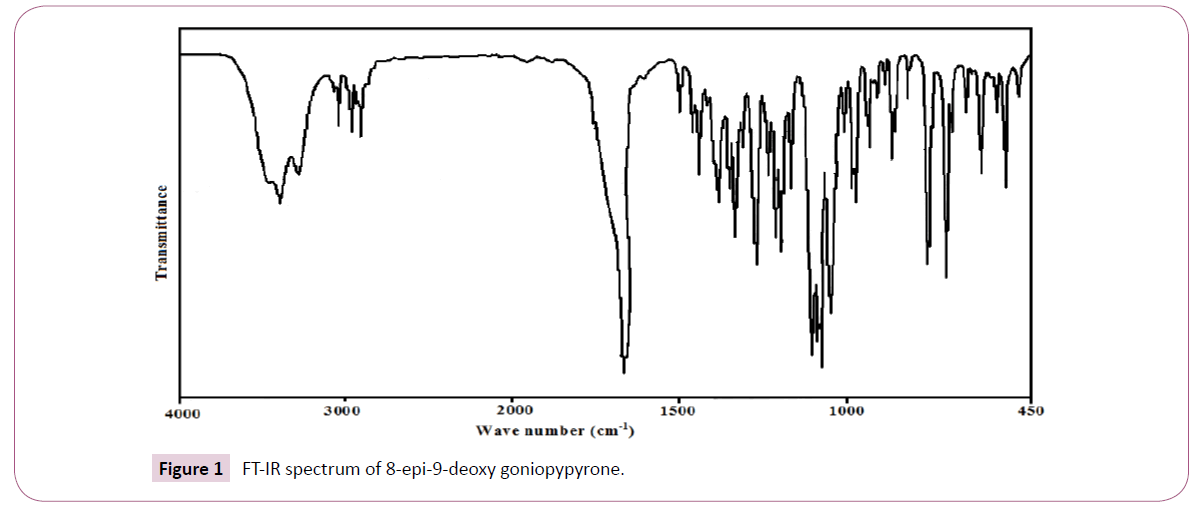
The plant material used in this study was collected from Peppara Wild Life Sanctury, Nedumangadu, Kerala, India in March 2004 and identified by Dr. N Mohan, Tropical Botanical Garden and Research Institute (TBGRI), Kerala, India. A voucher specimen (No.01998) has been deposited at the Herbarium of TBGRI, Kerala, India. Air-dried and finely powdered leaf (640 g) was soxlet extracted with hexane for 24 hours. The hexane extract was concentrated under reduced pressure and was kept open for two days at room temperature for the removal of traces of solvent. The concentrated viscous material (36.8 g) was chromatographed over a silicagel (100-200 mesh) column (1.5 m × 30 mm). The column was eluted gradually with hexane, mixture of hexane and chloroform and finally with pure chloroform. Aliquots of volume 100 ml were collected and each was checked by TLC. Fractions showing same TLC profiles were grouped together. The compound 8-epi-9-deoxy goniopypyrone was obtained as colorless crystals from hexane:chloroform (1:1 v/v). Recrystallization from the chloroform:methanol (1:1) yielded colourless crystals. The compound was identified by spectral analysis and by comparing the data with those reported in the literature. The compound 8-epi-9-deoxygoniopypyrone was previously isolated from Goniothalamus giganteus [10] and Goniothalamus leiocarpus [9]. The FT-IR spectrum (Figure 1) was recorded using KBr pellets on a DR/Jasco FT-IR spectrometer.
Computational details
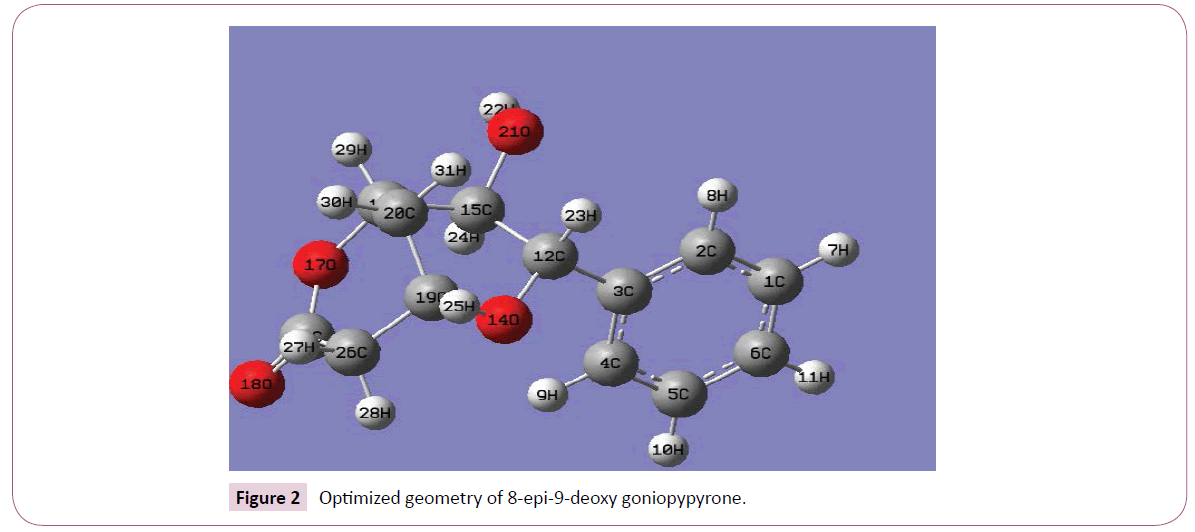
Calculations of the title compound were carried out with Gaussian09 software [11] program using HF/6-31G(6D,7F), B3LYP/6-31G(6D,7F) and B3LYP/311++G(5D,7F) basis sets to predict the molecular structure and vibrational wave numbers. Calculations were carried out with Becke’s three parameter hybrid model using the Lee–Yang–Parr correlation functional (B3LYP) method. Molecular geometry was fully optimized by Berny’s optimization algorithm using redundant internal coordinates. Harmonic vibrational wave numbers were calculated using analytic second derivatives to confirm the convergence to minima on the potential surface. Then frequency calculations were employed to confirm the structure as minimum points in energy. At the optimized structure (Figure 2) of the examined species, no imaginary wave number modes were obtained, proving that a true minimum on the potential surface was found. The DFT method tends to overestimate the fundamental modes; therefore scaling factor (0.9613) has to be used for obtaining a considerably bsetter agreement with experimental data [12]. The observed disagreement between theory and experiment could be a consequence of the anharmonicity and of the general tendency of the quantum chemical methods to overestimate the force constants at the exact equilibrium geometry. The assignments of the calculated wave numbers are aided by the animation option of GAUSSVIEW program [13] and potential energy distribution [14].
Results and Discussion
IR spectrum
The observed IR bands and calculated (scaled) wave numbers and assignments are given in (Table 1). The phenyl ring and pyrone ring are designated Ph and Py in the following discussion. The phenyl C-H stretching occurs above 3000 cm-1 and is typically exhibited as a multiplicity of weak to moderate bands, compared with the aliphatic C-H stretch [15]. The bands observed at 3077, 3066 cm-1 in the IR spectrum is assigned as the C-H stretching modes of the phenyl rings. The DFT calculations give these modes in the range 3075-3033 cm-1. The benzene ring possesses six ring stretching vibrations, of which the four with the highest wave numbers (occurring near 1600, 1580, 1490 and 1440 cm-1) are good group vibrations. In the absence of ring conjugation, the band near 1580 cm-1 is usually weaker than that at 1600 cm-1. The fifth ring stretching vibration, which is active near 1335 ± 35 cm-1 a region which overlaps strongly with that of the CH in-plane deformation [16]. The sixth ring stretching vibration or ring breathing mode appears as a weak band near 1000 cm-1 in mono, 1,3-di and 1,3,5-trisubstituted benzenes. In the otherwise substituted benzene, however, this vibration is substituent sensitive and difficult to distinguish from the ring in-plane deformation. For mono substituted benzene, the stretching modes are expected in the range of 1285-1610 cm-1 [16]. For the title compound, the stretching modes of phenyl ring are assigned at 1580, 1560, 1481, 1440, 1317 cm-1 theoretically and bands are observed at 1490, 1444 cm-1 in IR spectrum. The ring breathing mode for mono substituted benzenes appears near 1000 cm-1 [16]. For the title compound, this is confirmed by the band at 1000 cm-1 in the IR spectrum, which finds support from the computational results 993 cm-1. For mono substituted benzene, the in-plane CH vibrations are expected in the range of 1015-1300 cm-1 [16]. For the title compound these bands are observed at 1271 cm-1 in the IR spectrum and the corresponding theoretical values (DFT) are 1272, 1169, 1163, 1069, 1015 cm-1. The C-H out-of-plane deformations γCH are observed between 1000 and 700 cm-1 [16]. In the present case the out-of-plane CH modes are observed at 974, 942, 925, 751 cm-1 in the IR spectrum, and the corresponding theoretical values are 972, 943, 922, 844, 748 cm-1.
The carbonyl group is contained in a large number of different classes of compounds, for which a strong absorption band due to the C=O stretching vibration is observed in the region [17] 1850– 1550 cm-1. If a carbonyl group is part of a conjugated system, then the wave number of the carbonyl stretching vibration decreases, the reason being that the double bond character of the C=O group is less due to the π-electron conjugation being localized. For the title compound, the υC=O mode is seen as a strong band at 1650 cm-1 in the IR and at 1640 cm-1 theoretically assigned as the stretching mode of C=O [18] reported the C=O stretching mode is at 1658 cm-1 in the IR spectrum, at 1660 cm-1 in the Raman spectrum and at 1675 cm-1 theoretically. The deformation bands of C=O group are also identified (Table 1). The C-O stretching vibration [16] absorbs in the range 1045 ± 45 cm-1 [19] reported that the C-O stretching mode at 1024 cm-1 theoretically. In the present case C-O stretching mode are observed at 1103, 1162 cm-1 in the IR spectrum and at 1107, 1025, 1176, 1161 cm-1 theoretically.
| HF/6-31G(6D,7F) | B3LYP/6-31G(6D, 7F) | B3LYP/311++G (5D, 7F) | IRυ(cm-1) | Assignmentsa | |||
|---|---|---|---|---|---|---|---|
| υ(cm-1) | IRI | υ(cm-1) | IRI | υ(cm-1) | IRI | ||
| 3675 | 73.43 | 3609 | 28.53 | 3556 | 27.83 | υOH(100) | |
| 3402 | 6.44 | 3104 | 4.62 | 3075 | 6.92 | 3077 | υCHPh(94) |
| 3024 | 28.92 | 3086 | 18.38 | 3064 | 31.22 | 3066 | υCHPh(94) |
| 3015 | 37.22 | 3078 | 31.34 | 3052 | 32.16 | υCHPh(97) | |
| 3005 | 11.05 | 3067 | 13.88 | 3042 | 5.59 | υCHPh(95) | |
| 2995 | 0.36 | 3058 | 0.22 | 3033 | 0.65 | υCHPh(93) | |
| 2954 | 6.38 | 3012 | 4.66 | 2987 | 2.51 | υCH2(99) | |
| 2938 | 40.38 | 2999 | 19.25 | 2983 | 18.24 | υCH2(98) | |
| 2936 | 25.43 | 2974 | 35.77 | 2974 | 17.79 | 2977 | υCHPy(87), υCH2(11) |
| 2908 | 34.30 | 2959 | 35.77 | 2950 | 12.58 | 2952 | υCH2(79), υCHPy(12) |
| 2904 | 72.84 | 2945 | 9.12 | 2935 | 54.10 | 2933 | υCH2(90) |
| 2891 | 15.98 | 2944 | 21.45 | 2922 | 20.01 | υCHPy(97) | |
| 2884 | 10.99 | 2921 | 55.98 | 2910 | 7.04 | 2911 | υCHPy(95) |
| 2875 | 13.84 | 2888 | 20.09 | 2904 | 12.76 | 2900 | υCHPy(98) |
| 1803 | 503.57 | 1784 | 346.97 | 1640 | 350.42 | 1650 | υC=O(83) |
| 1620 | 2.85 | 1602 | 4.10 | 1580 | 2.85 | υPh(61), δPh(10), δCHPh(13) |
|
| 1595 | 0.99 | 1581 | 0.89 | 1560 | 0.81 | υPh(69) | |
| 1494 | 12.37 | 1487 | 12.41 | 1481 | 11.45 | 1490 | δCHPh(22), υPh(63) |
| 1479 | 16.22 | 1466 | 15.68 | 1466 | 18.12 | 1462 | δCH2(92) |
| 1449 | 12.38 | 1442 | 8.32 | 1440 | 8.12 | 1444 | δCHPh(16), υPh(59) |
| 1436 | 11.89 | 1426 | 7.59 | 1428 | 12.83 | 1426 | δCH2(93) |
| 1419 | 3.31 | 1387 | 5.68 | 1370 | 1.12 | 1375 | δOH(42), δCHPy(45) |
| 1395 | 44.49 | 1365 | 23.32 | 1357 | 2.05 | υPy(54) | |
| 1379 | 41.84 | 1352 | 7.01 | 1348 | 4.49 | 1350 | δCH2(40), δPy(27), δCHPh(10) |
| 1366 | 77.02 | 1343 | 34.69 | 1342 | 3.82 | δCHPy(22), δCH2(43), δPy(24) |
|
| 1357 | 17.98 | 1334 | 37.14 | 1331 | 19.98 | 1335 | δCHPh(18), δCH2(26), δPy(22) |
| 1345 | 20.08 | 1314 | 9.61 | 1317 | 6.13 | υPh(53), δCHPh(20) |
|
| 1333 | 5.34 | 1312 | 7.07 | 1303 | 14.44 | 1306 | δCH2(48), υCCPy(12) |
| 1322 | 14.52 | 1304 | 1.28 | 1293 | 6.07 | δPy(30), δCHPy(16) |
|
| 1313 | 19.36 | 1299 | 11.99 | 1285 | 7.18 | δPy(54), υPh(11) |
|
| 1302 | 16.46 | 1288 | 9.80 | 1272 | 16.83 | 1271 | δPy(14), δCHPh(48) |
| 1300 | 30.81 | 1275 | 19.97 | 1264 | 11.84 | υCCPy(24), δCH2(18) |
|
| 1220 | 145.05 | 1251 | 30.68 | 1233 | 35.99 | 1233 | δCH2(50) |
| 1210 | 51.55 | 1200 | 31.11 | 1203 | 38.60 | 1205 | υCCPy(49), δOH(11) |
| 1204 | 92.67 | 1190 | 66.58 | 1196 | 6.13 | 1195 | δOH(17), δCHPy(10), δCHPh(14), υCC(11) |
| 1191 | 5.04 | 1181 | 51.25 | 1176 | 1.95 | δCH2(23), υCO(46), υCO(12) |
|
| 1177 | 32.35 | 1169 | 35.83 | 1169 | 2.38 | δCHPh(74) | |
| 1164 | 1.11 | 1165 | 1.80 | 1163 | 0.09 | δCHPh(77), υPh(15) |
|
| 1142 | 48.80 | 1147 | 0.08 | 1161 | 54.37 | 1162 | υCO(47), δCCPy(16), δCH2(21) |
| 1127 | 273.42 | 1120 | 52.52 | 1107 | 43.39 | 1103 | υCO(44) |
| 1098 | 26.13 | 1084 | 175.46 | 1077 | 31.92 | 1075 | υCO(15), υCCPy(43), υPh(12) |
| 1093 | 51.28 | 1070 | 65.20 | 1069 | 2.84 | δCHPh(48), υCO(16) |
|
| 1076 | 46.52 | 1065 | 85.78 | 1043 | 77.80 | 1045 | υCO(H)(48) |
| 1069 | 1.87 | 1049 | 8.01 | 1025 | 217.37 | υCO(44), δCO(H)(13) |
|
| 1057 | 7.04 | 1042 | 43.21 | 1015 | 4.74 | δCHPh(40), υCCPy(15), tPy(16) |
|
| 1046 | 37.30 | 1030 | 15.96 | 993 | 48.09 | 1000 | υPh(53), δCHPh(12) |
| 1012 | 6.95 | 1018 | 10.86 | 989 | 9.42 | υCCPy(42), τPy(28) |
|
| 1006 | 0.84 | 987 | 2.73 | 984 | 6.55 | 982 | δPh(66), υPh(19) |
| 998 | 4.07 | 979 | 0.89 | 979 | 9.99 | υCO(16), υCCPy(26) |
|
| 992 | 17.11 | 970 | 42.29 | 972 | 2.84 | 974 | γCHPh(79), τPh(14) |
| 986 | 0.41 | 958 | 0.37 | 967 | 15.76 | δCH2(42), υCO(14) |
|
| 973 | 0.54 | 936 | 14.42 | 943 | 70.47 | 942 | γCHPh(86) |
| 960 | 23.68 | 933 | 1.02 | 922 | 14.40 | 925 | γCHPh(42), tPy(10), υCCPy(20) |
| 933 | 3.64 | 906 | 24.21 | 912 | 26.83 | 910 | δCH2(49), tPy(17) |
| 914 | 28.23 | 899 | 4.87 | 890 | 43.44 | 888 | γCHPh(49) |
| 911 | 7.36 | 889 | 4.97 | 879 | 8.28 | 872 | υCCPy(25), υCO(10), δCH2(10) |
| 873 | 4.80 | 865 | 6.72 | 847 | 0.38 | υCCPy(49), tPy(11) |
|
| 858 | 0.08 | 833 | 8.84 | 844 | 5.75 | γCHPh(99) | |
| 833 | 9.94 | 829 | 0.20 | 826 | 23.48 | 820 | υCCPy(41), υCO(13) |
| 798 | 5.14 | 795 | 6.40 | 778 | 12.78 | υCCPy(47), υCO(34) |
|
| 761 | 12.25 | 748 | 6.50 | 748 | 44.77 | 751 | γCHPh(48), tPh(30), γCC(10) |
| 756 | 30.41 | 739 | 24.17 | 716 | 5.05 | γCHPh(29), tPh(62) |
|
| 695 | 29.93 | 687 | 23.65 | 695 | 42.01 | 688 | δPh(25), δPy(34) |
| 648 | 6.93 | 642 | 4.79 | 645 | 4.69 | 642 | δPh(71) |
| 611 | 4.51 | 611 | 0.21 | 625 | 0.15 | 620 | δPy(21), δPh(26) |
| 606 | 1.67 | 604 | 1.80 | 599 | 4.02 | 597 | δPh(10), δPy(25) |
| 581 | 16.59 | 575 | 10.35 | 566 | 18.52 | 564 | γC=O(34), tPy(45) |
| 567 | 16.31 | 549 | 4.62 | 544 | 6.93 | 545 | δPy(23), υCCPy(12) |
| 529 | 14.06 | 526 | 11.61 | 526 | 9.74 | 524 | tPh(41), γCC(22) |
| 523 | 10.46 | 515 | 8.03 | 516 | 12.80 | 508 | δC=O(29), δPy(30) |
| 488 | 7.21 | 483 | 5.07 | 476 | 7.31 | 478 | δPy(43), δC=O(13), tPy(15) |
| 445 | 3.73 | 439 | 2.72 | 428 | 2.96 | δPy(18), tPy(11) |
|
| 414 | 0.68 | 408 | 1.63 | 406 | 1.10 | tPh(82) | |
| 406 | 0.15 | 399 | 0.24 | 404 | 1.14 | δPy(60), tPy(11) |
|
| 393 | 4.82 | 390 | 3.21 | 382 | 5.15 | tPy(15), tPh(20), δPy(19), δCO(13) |
|
| 374 | 11.12 | 367 | 9.29 | 362 | 4.84 | tOH(29) | |
| 295 | 26.85 | 298 | 45.67 | 284 | 0.91 | tOH(63) | |
| 274 | 51.36 | 280 | 65.14 | 265 | 2.95 | δCC(21), δPy(18), tPy(21) |
|
| 273 | 57.14 | 277 | 4.51 | 260 | 14.52 | tPy(70) | |
| 268 | 13.56 | 269 | 3.63 | 223 | 2.03 | tPy(45), tPh(12) |
|
| 223 | 0.04 | 222 | 0.13 | 205 | 113.80 | tPy(86) | |
| 204 | 3.05 | 199 | 2.25 | 193 | 36.32 | tPy(95) | |
| 107 | 2.73 | 106 | 1.99 | 101 | 3.31 | tPy(71) | |
| 87 | 0.71 | 87 | 0.83 | 81 | 0.77 | tPy(93) | |
| 73 | 2.27 | 74 | 1.97 | 73 | 2.92 | tPy(92) | |
| 40 | 3.26 | 39 | 2.28 | 37 | 3.75 | tPy(76) | |
| 30 | 0.41 | 30 | 0.44 | 24 | 0.38 | tCC(68), tPy(18) |
|
Table 1: Vibrational assignments of 8-epi-9-deoxygoniopypyrone.
The vibrations of the CH2 group, the asymmetric stretch, symmetric stretch, scissoring vibration and wagging vibration appear in the regions 3000 ± 50, 2965 ± 30, 1455 ± 55 and 1350 ± 85 cm-1, respectively [16-20]. For the title compound, DFT calculations give υasCH2 at 2987, 2983 cm-1 and υsCH2 at 2950, 2935 cm-1. Experimentally these bands are observed at 2952, 2933 cm-1 in the IR spectrum. For the title compound, DFT calculation gives bands at 1466, 1428 cm-1 as the scissoring mode δCH2 and these modes are observed at 1462, 1426 in the IR spectrum. The CH2 wagging modes are observed at 1350 cm-1 in IR spectrum and at 1348, 1342 cm-1 theoretically. The bands at 1306, 1233 cm-1 in IR spectrum and 1303, 1233 cm-1 (DFT) are assigned as the twisting mode τCH2. The rocking mode [16] is expected in the range 895 ± 85 cm-1. The band at 910 cm-1 (IR) and 967, 912 cm-1 (DFT) is assigned as the rocking CH2 mode for the title compound. The torsion modes of CH2 are seen in the low wave number range [16].
For the title compound stretching modes of OH bands are at 3556 cm-1 theoretically. The in-plane-deformation δOH is expected in the region 1400 ± 40 cm-1 [16]. In the present case the inplane OH bands are observed at 1375 cm-1 in the IR spectrum and at 1370 cm-1 theoretically. The C-O stretching mode (C-O) H band is observed at 1045 cm-1 experimentally and at 1043 cm-1 theoretically. The C-C stretching mode absorbs weakly to moderately in the region 915 ± 65 cm-1 [16].
For the title compound, the stretching bands of pyrone ring are observed at 1205, 1075, 820 cm-1 (IR) and at 1264, 1203, 1077, 989, 847, 826, 778 cm-1 (DFT). Sumayya et al. [21] reported the C-C stretching modes at 1082-900 (IR), at 1068-899 cm-1(HF). Pyrone ring stretching modes are reported at 1073, 1008, 880 cm-1 in the IR spectrum, at 1070, 1011, 881 cm-1 in the Raman spectrum and at 1068, 1000, 937, 905 cm-1 theoretically [21,22] reported the pyrone ring stretching modes at 1622, 1231, 910, 834 cm-1 in the IR spectrum, 1621, 1218, 1055, 895 cm-1 in the Raman spectrum and 1612, 1227, 1056, 900, 838 cm-1 theoretically. The CH stretching vibration in the pyrone ring is expected above 3000 cm-1 [16]. For the title compound these modes are observed at 2977, 2911, 2900 cm-1 and at 2974, 2922, 2910, 2904 cm-1 theoretically. For a pyrone derivative, the in-plane CH deformation modes are reported at 1403 cm-1 (IR), 1236 cm-1 (Raman), 1400, 1240 cm-1 (DFT) and the out-of-plane CH deformation modes are at 995 cm-1 (IR), 990 cm-1 (Raman), 999, 825 cm-1 (DFT) [22]. In the present study these bands are observed at 1375 cm-1 in the IR spectrum and the corresponding DFT values are in the range 1370-1285 cm-1. The CH modes are calculated theoretically in the range 984-879 cm-1 and are assigned in the (Table 1). Most of the modes are not pure but contains significant contribution from other modes also. The RMS error of the observed IR bands is 57.37, 21.97 and 3.93 for HF/6-31G(6D,7F), B3LYP/6-31(6D,7F) and B3LYP/311++G(5D,7F) methods.
Natural bond analysis
The natural bond orbitals (NBO) calculations were performed using NBO 3.1 program [23] as implemented in the Gaussian09 package at the DFT/B3LYP level in order to understand various second-order interactions between the filled orbitals of one subsystem and vacant orbitals of another subsystem, which is a measure of the intra-molecular delocalization or hyper conjugation. The second-order perturbation theory analysis of Fock-matrix in NBO basis shows strong intra-molecular hyper conjugative interactions are formed by orbital overlap between n(O) and σ*(C-C), σ*(C-O), π*(C-O) bond orbitals which result in intra-molecular charge transfer causing stabilization of the system. There occurs an intra-molecular hyper conjugative interaction C12-C15 from O14 of n2(O14)→σ*(C12-C15) which increases ED (0.04626e) and weakens the respective bonds C12-C15 leading to stabilization of 6.47 KJ/mol and a strong intra-molecular hyper conjugative interaction C13-O18 from O17 of n2(O17)→π*(C13-O18) which increases ED (0.19751e) and weakens the respective bonds C13-O18 leading to stabilization of 39.12 KJ/mol. Another hyper conjugative interaction of C13-O17 from O18 of n2(O18)→σ*( C13-O17) which increases ED (0.11095e) and weakens the respective bonds C13-O17 leading to stabilization of 35.84 KJ/mol. There occurs an intra-molecular hyper conjugative interaction of C15-C16 from O21 of n2(O21)→σ*(C15-C16) which increases ED (0.04921e) and weakens the respective bonds C15-C16 leading to stabilization of 8.45 KJ/mol. These interactions are observed as an increase in electron density in C-C anti-bonding orbital that weakens the respective bonds. The hyper conjugative interaction energy was deduced from the second-order perturbation approach. Delocalization of electron density between occupied Lewistype (bond or lone pair) NBO orbitals and formally unoccupied (anti bond or Rydberg) non-Lewis NBO orbitals corresponds to a stabilizing donor-acceptor interaction.
The NBO analysis describes the bonding in terms of the natural hybrid orbital n2(O14), which occupy a higher energy orbital (-0.29945 a.u.) with considerable p-character (99.96%) and low occupation number (1.92418) and the other n1(O14) occupy a lower energy orbital (-0.55556 a.u.) with p-character (58.38%) and high occupation number (1.96164). The NBO analysis also describes the bonding in terms of the natural hybrid orbital n2(O17), which occupy a higher energy orbital (-0.31775 a.u.) with considerable p-character (99.86%) and low occupation number (1.79762) and the other n1(O17) occupy a lower energy orbital (-0.53194 a.u.) with p-character (42.70%) and high occupation number (1.95647). The NBO analysis also describes the bonding in terms of the natural hybrid orbital n2(O18), which occupy a higher energy orbital (-0.24356 a.u.) with considerable p-character (99.93%) and low occupation number (1.83803) and the other n1(O18) occupy a lower energy orbital (-0.66886 a.u.) with p-character (42.70%) and high occupation number (1.97771). The NBO analysis describes the bonding in terms of the natural hybrid orbital n2(O21), which occupy a higher energy orbital (-0.31300 a.u.) with considerable p-character (99.48%) and low occupation number (1.95062) and the other n1(O21) occupy a lower energy orbital (-0.60958 a.u.) with p-character (51.50%) and high occupation number (1.98175). Thus, a very close to pure p-type lone pair orbital participates in the electron donation to the n2(O14)→σ*(C12-C15), n2(O17)→π*(C13-O18), n2(O18)→σ*( C13-O17) and n2(O21)→σ*(C15-C16) interactions in the compound.
Frontier molecular orbital analysis
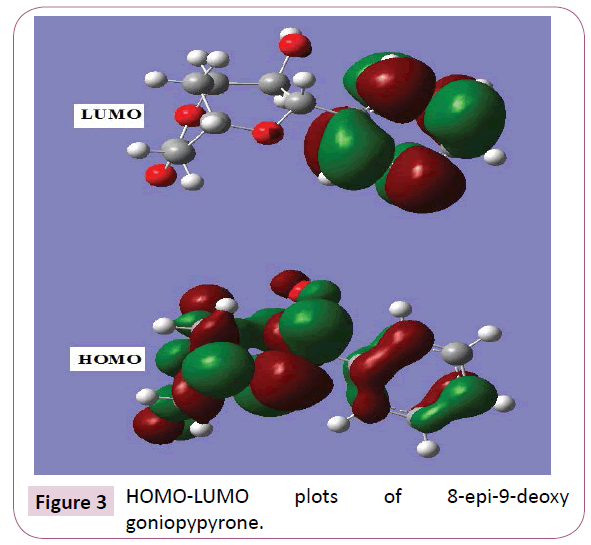
The most widely used theory by chemists is the molecular orbital (MO) theory. It is important that ionization potential (I), electron affinity (A), electrophilicity index (ω), chemical potential (μ), electronegativity (χ) and hardness (η) to be put into a molecular orbital frame work. Based on density functional descriptors, global chemical reactivity descriptors of compounds such as hardness, chemical potential, softness, electro negativity and electrophilicity index as well as local reactivity has been defined [24]. Pauling introduced the concept of electronegativity as the power of an atom in a compound to attract electrons to it. Using Koopman’s theorem for closed shell components η, μ and χ can be defined as η = (I -A)/2; μ = -(I + A)/2; χ = (I + A)/2; where I and A are the ionization potential and electron affinity of the compounds respectively. The ionization energy (I) and electron affinity (A) can be expressed through HOMO and LUMO orbital energies as I=-EHOMO and A=-ELUMO. Electron affinity refers to the capability of ligand to accept precisely one electron from a donor. However, in many kinds of bonding viz. covalent hydrogen bonding, partial charge transfer takes place. Considering the chemical hardness (η), large HOMO-LUMO energy gap means a hard molecule and small HOMO-LUMO gap means a soft molecule. One can also relate the stability of the molecule to hardness, which means that the molecule with smaller HOMO-LUMO gap is more reactive [24] have defined a new descriptor to quantity the global electrophilic power of the compound as electrophilicity index (ω) which defines a quantitative classification of global electrophilic nature of a compound. Parr et al. have proposed electrophilicity index (ω) as a measure of energy lowering due to maximal electron flow between donor and acceptor. They defined electrophilicity index as follows: ω=μ2/2η. The usefulness of this new reactivity measure has been recently demonstrated in understanding the toxicity of various pollutants in terms of their reactivity and site selectivity [25]. The calculated values of ω, μ, χ and are 12.2387 eV, -7.3439 eV, 7.3439 eV and 2.2039 eV respectively. The calculated value of electrophilicity index describes the biological activity of the title compound. The atomic orbital components of the frontier molecular orbital are shown in Figure 3.
Molecular electrostatic potential
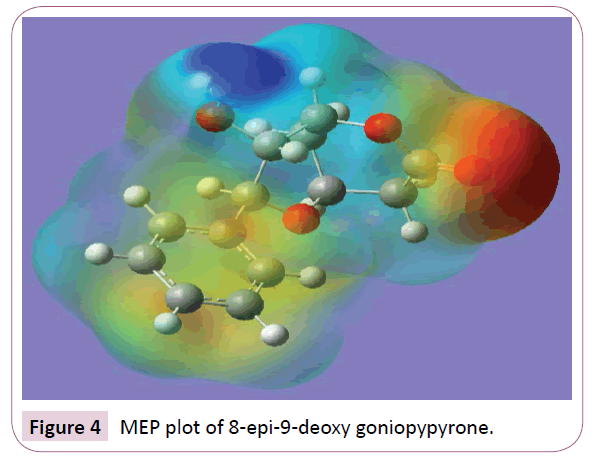
Molecular electrostatic potential (MEP) is a charge distribution in the space around the molecule. MEP is very useful descriptor in understanding sites for electrophilic attack and nucleophilic reactions and for the study of biological recognition process [26,27]. To predict reactive sites of electrophilic and nucleophilic attacks for the investigated molecule, MEP at the B3LYP/6-31G (6D,7F) optimized geometry was calculated. Figure 4 provides a visual presentation of the chemically active sites and comparative reactivity of atoms. The potential has been particularly useful as an indicator of the sites or regions of a molecule which are initially attracted by approaching electrophile/nucleophile, and it has also been successfully applied to determine optimum relative orientation of the reactants. Potential value increases in the order red<orange<yellow<green<blue. The negative electrostatic potential corresponds to an attraction of the proton by the aggregate electron density in the molecule (shades of red), while the positive electrostatic potential corresponds to the repulsion of the proton by the atomic nuclei (shade of blue). The negative (red and yellow) regions of MEP were related to electrophilic reactivity and the positive (blue) regions to nucleophilic reactivity. From the MEP of the title compound it is evident that the negative charge covers the C=O group. The positive region is over the hydroxyl group. The value of the electrostatic potential is largely responsible for the binding of a substrate to its receptor binding sites since the receptor and the corresponding ligand recognize each other at their molecular surface [28].
Nonlinear optical properties
Nonlinear optics deals with the interaction of applied electromagnetic fields in various materials to generate new electromagnetic fields, altered in wave number, phase or other physical properties. Organic molecules able to manipulate photonic signals efficiently are of importance in technologies such as optical communication, optical computing and dynamic image processing [29,30]. The calculated first hyperpolarizability of the title compound is 0.6369 × 10-30 e.s.u. which is 5 times that of standard NLO material urea (0.13 × 10-30 e.s.u.) [31]. The second hyperpolarizability is calculated using the using the following formula.
γav= [γ xxxx + γ yyyy + γ zzzz + 2 γ xxyy + 2 γ xxzz + 2 γ yyzz]/5. The second hyperpolarizability of the title compound is -9.016 × 10-37 e.s.u [32]. Thus the present material has a reasonably good reactivity for nonlinear optical activity.
Molecular docking
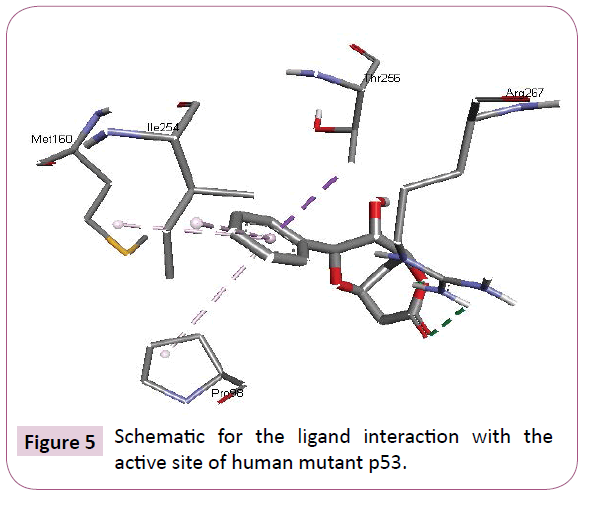
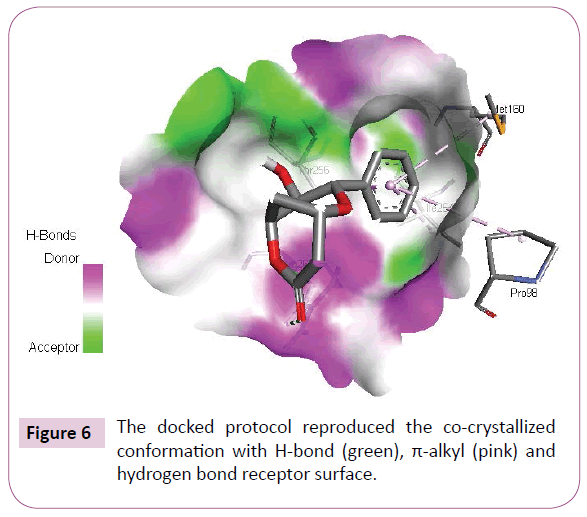
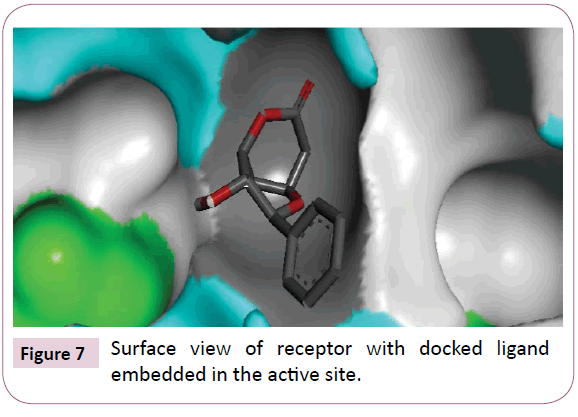
Acetogenins and styryl-lactones from Goniothalamus species have shown to be cytotoxic to different human tumour cell lines [33,34]. Styryllactones of Goniothalamusgitin genesis (goniopypyrone) are active against Mycobacterium tuberculosis H37Rv and cancer cell lines [35,36]. The constituent of ethyl acetate extract of G. marcanii contain three derivative styryllactones such as 5-hydroxygoniothalamin, 5-acetylgoniothalamin and goniopypyrone. These extract against human tumour cell lines including lung carcinoma (A549), colonadenocarcinoma (HT-29), breast carcinoma (MCF-7), melanoma (RPMI) and brain carcinoma (U251) [37]. The crude hexane extract of G. marcaniiexhibited cytotoxicity against MCF-7, murine lymphocytic leukemia (P-388), human oral nasopharyngeal carcinoma (KB), human colon cancer (Col-2), human lung cancer (Lu-1), rat glioma (ASK), noncancerous human embryonic kidney (Hek 239) and human urinary bladder (T24) cell lines [38]. High resolution crystal structure of human mutant p53 was downloaded from the RSCB protein data bank website (PDB ID: 4MZI). All molecular docking calculations were performed on Auto Dock Vina software [39]. The 3D crystal structure of human mutant p53 was obtained from Protein Data Bank. The protein was prepared for docking by removing the cocrystallized ligand, waters and co-factors. The Auto Dock Tools (ADT) graphical user interface was used to calculate Kollman charges and polar hydrogen’s. The ligand was prepared for docking by minimizing its energy at B3LYP/6-31G (6D, 7F) level of theory. Partial charges were calculated by Geistenger method. The active site of the enzyme was defined to include residues of the active site within the grid size of 40Å × 40Å × 40Å. The most popular algorithm, Lamarckian Genetic Algorithm (LGA) available in Autodock was employed for docking. The docking protocol was tested by extracting co-crystallized inhibitor from the protein and then docking the same. The docking protocol predicted the same conformation as was present in the crystal structure with RMSD value well within the reliable range of 2Å [40]. Amongst the docked conformations, one which binds well at the active site was analyzed for detailed interactions in Discover Studio Visualizer 4.0 software. The ligand binds at the active site of the substrate (Figures 5 and 6) by weak non-covalent interactions. Arg267 amino acid forms H-bond interaction with C=O and pyrone ring. Amino acids Pro98, Ile254, Thr256 and Met160 forms π-alkyl interaction with phenyl ring. Thr256 amino acid form π-sigma interactions with phenyl ring. The docked ligand title compound forms a stable complex with human mutant p53 and gives a binding affinity (ΔG in kcal/mol) value of -6.1 (Table 2). These preliminary results suggest that the compound might exhibit inhibitory activity against human mutant p53 (Figure 7).
| Mode | Affinity (kcal/mol) | Distance from best mode (Å) | |
|---|---|---|---|
| RMSD l.b. | RMSD u.b. | ||
| 1 | -6.1 | 0.000 | 0.000 |
| 2 | -6.1 | 18.695 | 20.852 |
| 3 | -5.7 | 14.469 | 16.164 |
| 4 | -5.6 | 2.363 | 3.089 |
| 5 | -5.6 | 3.295 | 5.067 |
| 6 | -5.5 | 18.637 | 21.010 |
| 7 | -5.5 | 20.611 | 22.120 |
| 8 | -5.4 | 17.573 | 19.426 |
| 9 | -5.4 | 3.480 | 4.208 |
Table 2: The binding affinity values of different poses of the title compound predicted by Autodock Vina.
Conclusion
The vibrational spectroscopic studies of the 8-epi-9-deoxy goniopypyrone in the ground state were reported experimentally and theoretically. The lowering of HOMO-LUMO band gap supports for the bioactivity of the molecule. From MEP, the most reactive part in the molecule is the C=O group. The calculated hyperpolarizability title compound is -9.016 × 10-37 esu and the title compound is an attractive object for future studies of nonlinear optical properties. The title compound forms a stable complex with human mutant p53 and gives a binding affinity value of -6.1 kcal/mol and the compound might exhibit inhibitory activity against human mutant p53.
Acknowledgaements
Authors, B. Harikumar would like to thank UGC, India for a minor research project and KS. Resmi would like thank University of Kerala for a research fellowship.
References
- Alipio BC, David FC, Isabel RS, Juan CNM. Manuel AE. Comparison of batch, stirred flow chamber, and column experiments to study adsorption, Desorption and transport of carbofuran within two acidic soils. Chemosphere. 2012; 88(1): 106-120.
- Arnaud B, Richard C, Michel S. A comparison of five pesticides adsorption and Desorption processes in thirteen contrasting field soils. Chemosphere. 2005; 61(5): 668-676.
- Chunxian W, Jin-Jun W, Su-Zhi Z, Zhong-Ming Z. Adsorption and Desorption of Methiopyrsulfuron in Soils. Pedosphere. 2011; 21(3): 380-388.
- Chunxian W, Suzhi Z, Guo N, Zhongming Z, Jinjun W. Adsorption and Desorption of herbicide monosulfuron-ester in Chinese soils. J Environ Sci. 2011; 23(9): 1524-1532.
- Christine MFB, Josette MF. Adsorption-desorption and leaching of phenylurea herbicides on soils. Talanta. 1996; 43(10): 1793-1802.
Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences